I. INTRODUCTION
Prostate cancer diagnosis represents a critical challenge in modern urological oncology, where accurate early detection directly correlates with treatment efficacy and patient survival rates [1]. Traditional diagnostic pathways rely heavily on a combination of serum prostate-specific antigen (PSA) measurements, digital rectal examination findings, and transrectal ultrasound-guided biopsy results [2]. However, these conventional methods exhibit notable limitations in sensitivity and specificity, frequently leading to overdiagnosis of indolent tumors or delayed detection of aggressive malignancies [3]. The integration of multimodal medical data—encompassing magnetic resonance imaging sequences, histopathological patterns, laboratory biomarkers, and clinical narratives—offers promising avenues for enhancing diagnostic precision. However, the heterogeneous nature of these data sources, combined with their complex interrelationships, poses substantial computational challenges for automated analysis systems.
Recent advances in deep learning have catalyzed significant progress in medical image analysis and clinical decision support [4-5]. Convolutional neural networks demonstrate remarkable capability in extracting hierarchical features from radiological images, while recurrent architectures excel at processing sequential clinical records [6]. Nevertheless, these data-driven approaches exhibit fundamental weaknesses when confronting medical diagnosis tasks [7]. Deep neural networks typically require extensive annotated datasets for adequate training, which remain scarce in specialized domains like prostate pathology [8]. Further-more, purely data-driven models lack mechanisms for incorporating decades of accumulated clinical expertise regarding disease manifestations, risk factors, and diagnostic criteria [9]. This absence of structured medical knowledge frequently results in models that capture superficial statistical patterns while missing crucial clinical relationships that experienced urologists intuitively recognize [10].
Medical knowledge graphs represent a paradigm shift in how clinical expertise can be formalized and computationally leveraged [11-12]. These structured knowledge bases encode entities such as diseases, symptoms, biomarkers, and anatomical structures, along with their semantic relationships, including causal links, diagnostic associations, and hierarchical taxonomies [13]. For prostate cancer specifically, knowledge graphs can capture intricate relationships between Gleason scoring patterns, PSA kinetics, imaging characteristics on T2-weighted and diffusion-weighted MRI sequences, and ultimate pathological diagnoses [14]. Several large-scale medical knowledge graphs have emerged, including specialized oncology knowledge bases that document evidence-based relationships between clinical findings and cancer subtypes [15]. Unlike traditional knowledge representation formats, graph-structured knowledge facilitates efficient reasoning and enables machine learning models to traverse relationship paths during inference [16].
The intersection of knowledge graphs with deep learning architectures presents compelling opportunities for advancing prostate cancer diagnosis [17-18]. By embedding structured clinical knowledge into continuous vector spaces, these entities can be integrated directly into neural network training pipelines [19]. Consider the diagnostic scenario where a patient presents with moderately elevated PSA levels, an enlarged prostate on imaging, and nocturia symptoms [20]. A purely data-driven model might focus predominantly on the PSA elevation, potentially leading to unnecessary biopsies [21]. However, an approach augmented with knowledge graph representations would recognize that these symptoms collectively align more strongly with benign prostatic hyperplasia rather than malignancy, particularly when certain imaging features are absent [22]. The knowledge graph captures that while PSA elevation occurs in both conditions, specific patterns of elevation combined with particular MRI characteristics differentiate the two entities.
Building upon these observations, the MP-KDNet framework is introduced for multimodal prostate cancer diagnosis that synergistically combines knowledge graph embeddings with convolutional neural architectures. The methodology extracts structured pathological knowledge from medical knowledge graphs through entity alignment and graph embedding techniques, transforming this knowledge into dense vector representations. These knowledge vectors are then fused with multimodal clinical feature representations—derived from MRI imaging data, laboratory measurements, and clinical documentation—to form multi-channel inputs for a specialized convolutional network. Through this knowledge-enhanced learning process, the model simultaneously learns from objective multimodal patient data and subjective clinical expertise encoded in knowledge structures.
The contribution of this paper can be summarized as follows.
-
(1) We propose MP-KDNet, a novel framework that synergistically integrates medical knowledge graphs with multimodal deep learning for prostate cancer diagnosis, addressing the limitations of purely data-driven approaches in capturing clinical expertise.
-
(2) We construct a comprehensive prostate cancer knowledge graph containing 25,264 entities and 70,049 triples, then employ TransD embedding to transform structured clinical knowledge into continuous representations compatible with neural processing.
-
(3) We design a knowledge-enhanced multi-channel convolutional architecture that simultaneously processes multimodal clinical features, aligned knowledge entity embeddings, and contextual relationship vectors through parallel information streams with multi-scale pattern detection.
-
(4) We develop entity alignment and contextual knowledge extraction mechanisms that automatically link clinical observations from MRI reports, laboratory measurements, and patient documentation to relevant knowledge graph entities and their semantic neighborhoods.
The rest of the paper is organized as follows: Section II establishes the foundation for knowledge integration by describing how the prostate cancer knowledge graph is constructed and represented. Section III presents the complete MP-KDNet framework, walking through each component of the diagnostic pipeline. Section IV provides experimental validation of the proposed approach. Section V concludes the paper.
II. PROSTATE CANCER KNOWLEDGE GRAPH CONSTRUCTION
A prostate cancer knowledge graph GPCa constitutes a structured representation of clinical entities and their semantic relationships within the prostate cancer domain. The graph encodes medical knowledge through a collection of triples, where each triple formalizes a specific clinical relationship. Formally, GPCa is defined as:
where ξh denotes the head entity, ξt represents the tail entity, and ρ ∈ {ρ1, ρ2, … ρ|R|} specifies the relationship type connecting them. Both ξh and ξt belong to the entity set ε of GPCa, while ρ belongs to the relationship set R containing |R| distinct relationship types.
The entity set ε encompasses diverse medical concepts relevant to prostate cancer diagnosis. Primary entities include pathological conditions (adenocarcinoma, prostatic intraepithelial neoplasia, benign prostatic hyperplasia), imaging biomarkers (PI-RADS scores, apparent diffusion coefficient values, T2-weighted signal characteristics), laboratory measurements (PSA levels, PSA density, free-to-total PSA ratio), anatomical structures (peripheral zone, transition zone, anterior fibromuscular stroma), clinical manifestations (lower urinary tract symptoms, hematuria, bone pain), and therapeutic interventions (active surveillance, radical prostatectomy, radiation therapy).
The relationship set R captures various semantic associations between entities. Key relationship types include diagnostic_indicator (linking imaging findings to pathological states), risk_factor (connecting predisposing conditions to cancer development), prognostic_marker (relating biomarkers to disease outcomes), anatomical_ location (specifying tumor spatial distribution), treatment_ response (associating therapies with clinical outcomes), and disease_progression (tracking temporal evolution of pathological states).
For instance, the clinical knowledge that "peripheral zone adenocarcinoma typically demonstrates restricted diffusion on DWI sequences with ADC values below 1.0 × 10−3 mm2/s translates into the triple representation. The comprehensive knowledge graph aggregates thousands of such triples extracted from clinical guidelines, peer-reviewed literature, and electronic health records to construct a holistic representation of prostate cancer domain knowledge.
Knowledge graph embedding transforms discrete symbolic entities and relationships from GPCa into continuous low-dimensional vector spaces while presserving the graph's semantic structure [23]. This vectorization enables downstream integration with neural network architectures [24]. Given the complexity of prostate cancer relationships—including one-to-many, many-to-one, and many-to-many associations—the TransD embedding model is adopted for its superior handling of heterogeneous relationship patterns.
For a triple <ξh, ρ, ξt>, TransD posits that entities connected through relationship ρ occupy distinct semantic spaces. The model employs projection matrices Mh and Mt to map head and tail entities into the relationship-specific space. These projection matrices are decomposed as:
where ρp ∈ ℝκ represents the relationship projection vector, ξhp, ξtp ∈ ℝδ denote entity-specific projection vectors, and Iδ×κ is the identity matrix of appropriate dimensions. The scoring function measuring triple plausibility becomes:
This formulation allows each entity-relationship pair to define unique projection dynamics, accommodating the diverse semantic characteristics present in medical knowledge. Training optimizes embeddings by minimizing ϕρ for valid triples while maximizing scores for corrupted negatives, yielding entity vectors ξh, ξt ∈ ℝκ and relationship vectors ρ ∈ ℝκ that encode structured clinical knowledge in continuous space.
III. MULTIMODAL PROSTATE CANCER DIAGNOSIS VIA KNOWLEDGE-ENHANCED NETWORKS
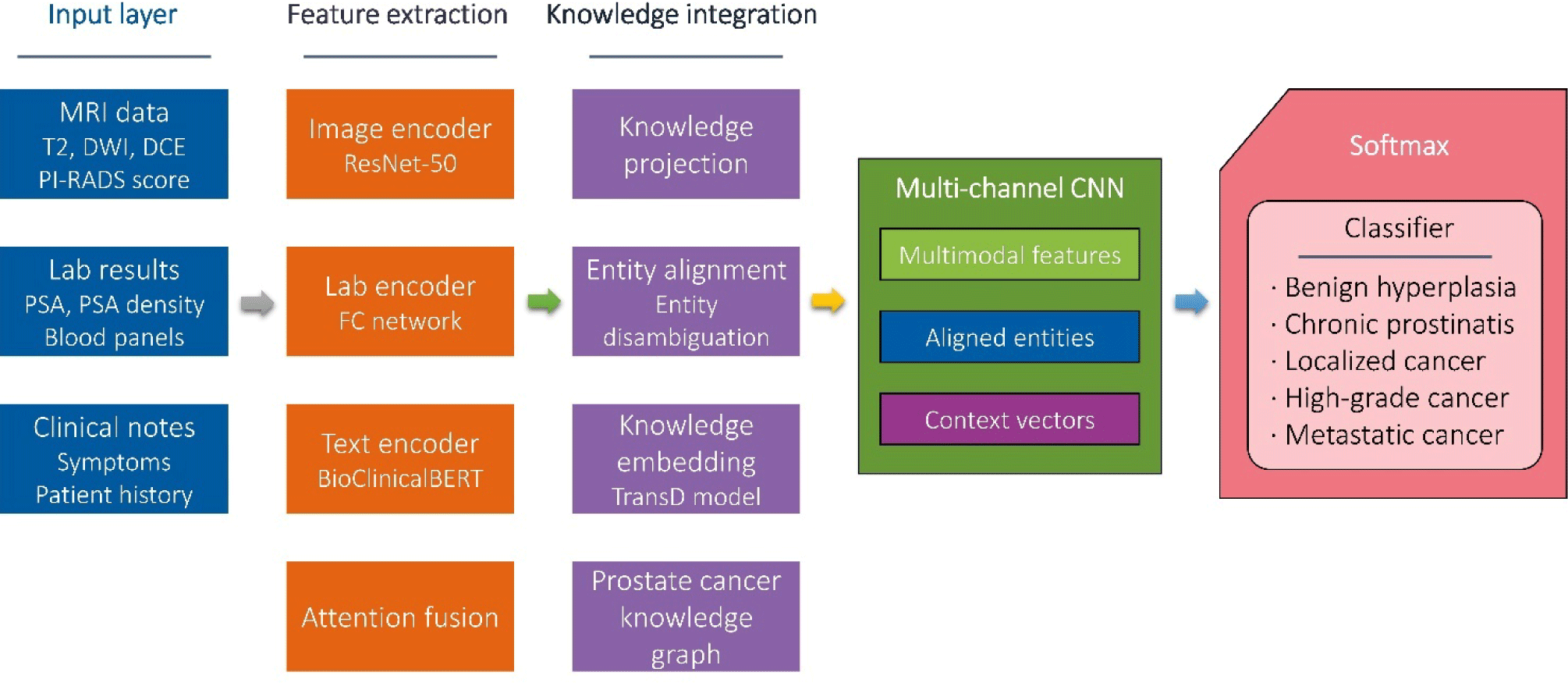
The MP-KDNet framework addresses prostate cancer diagnosis through three synergistic components: multimodal feature extraction, structured knowledge integration, and knowledge-enhanced convolutional classification. Fig. 1 illustrates the complete architecture, demonstrating information flow from raw multimodal inputs through knowledge-augmented processing to diagnostic outputs.
The input layer accepts heterogeneous clinical data sources: (1) MRI sequences including T2-weighted, diffusion-weighted, and dynamic contrast-enhanced imaging; (2) laboratory measurements encompassing PSA kinetics, complete blood counts, and metabolic panels; (3) clinical documentation comprising patient histories, symptom descriptions, and physical examination findings. These diverse modalities undergo specialized preprocessing pipelines tailored to their respective data types. Missing data were addressed through domain-informed imputation. For imaging data, missing sequences were imputed using zero-padding with a missing indicator channel. Laboratory values were imputed using cohort-specific median values stratified by age and diagnostic category. Text data missing specific sections was processed with available portions only. We evaluated model performance under systematic missing data scenarios, where randomly excluding one modality reduced accuracy by an average of 3.2 percentage points, demonstrating reasonable robustness to incomplete records.
The knowledge integration module establishes connections between extracted clinical features and entities within GPCa. Through entity alignment mechanisms, clinical descriptors from multimodal inputs are mapped to corresponding knowledge graph entities, enabling the retrieval of relevant structural knowledge. Knowledge graph embeddings transform these discrete entities into continuous representations that can be processed alongside data-driven features.
The convolutional architecture receives multi-channel inputs combining data-derived feature vectors with knowledge entity embeddings. Multiple convolutional kernels of varying receptive fields capture patterns across both modality-specific features and cross-modal interactions. The knowledge-enhanced representations enable the network to recognize diagnostically significant patterns that might be overlooked by purely data-driven approaches, particularly those involving subtle combinations of findings that experienced clinicians learn to identify through years of practice.
Clinical deployment would position MP-KDNet as a decision support tool rather than an autonomous diagnostic system. The model could process preliminary MRI readings, laboratory results, and clinical notes to generate diagnostic hypotheses before formal multidisciplinary tumor board review. Output probabilities across the five diagnostic categories would flag cases requiring urgent attention (high probability of aggressive cancer) versus those appropriate for conservative management (high probability of benign conditions). Attention visualizations showing influential features could guide specialists toward relevant findings warranting detailed examination. However, final diagnostic and treatment decisions would remain with treating physicians who integrate model outputs with additional clinical judgment and patient preferences.
Clinical presentation of prostate pathology manifests across multiple data modalities, each providing complementary diagnostic information. Effective multimodal fusion requires extracting semantically meaningful representations from each modality before integration.
For MRI imaging data, pre-trained ResNet architectures extract spatial features from each sequence type [25]. Given an MRI volume V ∈ ℝH×W×D with spatial dimensions H × W × D, the imaging encoder Φimg produces a feature map:
where dimg denotes the imaging feature dimensionality. Separate encoders process T2-weighted, DWI, and DCE sequences, with features subsequently concatenated to form a comprehensive imaging representation.
Laboratory measurements form a structured vector vlab ∈ ℝnlab containing nlab distinct biomarkers. These continuous values undergo normalization and dimensionality expansion through a fully-connected network:
where Wlab ∈ ℝdlab × nlab represents learnable weights, blab ∈ ℝdlab denotes bias terms, and σ(·) applies a nonlinear activation function.
Clinical text documents require specialized natural language processing [26]. Utilizing BioClinicalBERT pretrained on medical corpora, raw text T transforms into contextualized embeddings [27]:
These textual features capture semantic nuances in symptom descriptions, medical histories, and examination findings that often contain subtle diagnostic clues.
We analyzed attention weight distributions across cases to assess modality weighting patterns. Attention weights varied considerably across patients, with imaging receiving a higher weight (mean 0.42) in cases with clear radiological findings, while text features dominated (mean 0.51) when imaging showed equivocal patterns but detailed symptom histories existed.
The multimodal representation aggregates features across modalities through learned attention-weighted fusion [28]:
where attention weights αm are computed via:
with wm ∈ ℝm representing modality-specific attention parameters. This adaptive weighting allows the model to emphasize more diagnostically informative modalities for individual cases.
Integrating structured medical knowledge with data-driven features requires establishing correspondences between clinical observations and knowledge graph entities, followed by the extraction of relevant graph neighborhoods.
Clinical features extracted from multimodal data often reference medical concepts that exist as entities in GPCa. For instance, textual mentions of "elevated PSA" or imaging-derived "PI-RADS score" correspond to specific knowledge graph entities. Entity linking establishes these mappings through similarity computation between feature representations and entity embeddings.
Given a clinical descriptor fc and candidate entity embeddings {ξ1, ξ2 …, ξK}, the alignment score for candidate ξk is computed as:
where T ∈ ℝdc×κ is a learned transformation matrix bridging the feature space and entity embedding space. The entity with maximum alignment score is selected:
Entity disambiguation handles ambiguous medical terms through context-aware matching. When a clinical mention matches multiple candidate entities, we compute contextual similarity by comparing the sentence-level BioClinical-BERT embedding of the mention with the embeddings of entity definitions and neighboring entities in the knowledge graph. For instance, 'mass' could refer to prostatic mass or body mass index, disambiguated by examining whether the surrounding text discusses imaging findings versus anthropometric measurements.
Unmatched clinical mentions occurred in approximately 15% of entity alignment attempts, typically involving colloquial symptom descriptions or rare clinical presentations not captured in the knowledge graph. For unmatched entities, we implemented a fallback strategy using the original text feature embedding without knowledge augmentation for that specific feature position. This approach prevents information loss from failed matches while maintaining the multi-channel architecture.
Individual entities provide limited information; diagnostic reasoning often involves considering relationships to connected entities. For each aligned entity ξ, its knowledge context C(ξ) comprises one-hop neighbors in GPCa:
This context captures entities directly related through any relationship type ρ. For example, if ξ represents "peripheral zone lesion," C(ξ) would include connected entities like "adenocarcinoma risk," "PI-RADS 4-5 likelihood," and "targeted biopsy indication."
One-hop neighborhoods were selected after comparing the extraction depths of one, two, and three hops on a validation subset. Two-hop neighborhoods increased context size by an average of 8.7x, introducing many weakly-related entities that diluted the diagnostic signal. One-hop neighborhoods capture immediately relevant relationships such as diagnostic_indicator and risk_factor while avoiding the noise from distant entities connected through multiple relationship chains.
The contextual representation aggregates neighbor embeddings through mean pooling:
where |C(ξ)| denotes the context cardinality. This averaged context vector ξ̅ ∈ ℝκ encodes supplementary knowledge beyond the isolated entity, enriching the model's understanding of clinical implications.
For a complete clinical case with multiple aligned entities , a case-specific knowledge subgraph Gsub is extracted by collecting all entities and relationships connecting them within GPCa. This subgraph provides a structured representation of how the patient's clinical features interrelate according to established medical knowledge.
The core diagnostic module employs a multi-channel convolutional network that processes both multimodal features and knowledge representations simultaneously. This architecture enables learning patterns that integrate empirical observations with structured medical expertise.
For a patient case, let f1, f2, …, fM denote M extracted multimodal feature vectors, where each fi ∈ ℝδ represents features at a specific anatomical location or temporal measurement. Corresponding knowledge entities with their contexts are retrieved through the alignment procedure.
Entity embeddings require transformation from the knowledge space ℝκ to the feature space ℝδ for compatibility with multimodal features. A learnable projection accomplishes this mapping:
where Wproj ∈ ℝδ×κ and bproj ∈ ℝδ are trainable para-meters. The hyperbolic tangent activation ensures the projected entities occupy a similar value range as normalized features.
Hyperbolic tangent activation was selected for knowledge projection based on empirical comparison with ReLU, Leaky ReLU, and GELU. Tanh restricts projected embeddings to the range (-1, 1), matching the normalized range of multimodal features after standardization, which facilitates stable multi-channel fusion. In contrast, ReLU produced asymmetric value distributions (zero for negative inputs) that created imbalanced channel contributions, reducing validation accuracy by 1.4 percentage points. GELU performed comparably to tanh but offered no clear advantage while adding computational overhead.
Similarly, context vectors transform as:
The multi-channel input tensor combines these three information sources:
Here, the three channels correspond to: (1) data-driven multimodal features, (2) aligned knowledge entities, and (3) entity contexts. This structure parallels RGB image channels, enabling standard convolutional operations.
While our convolutional architecture captures local patterns across feature vectors, it processes observations as an unordered set rather than explicitly modeling spatial or temporal structure. For imaging features extracted from different prostate zones, spatial adjacency information is preserved through the ResNet encoder but not explicitly leveraged during knowledge-enhanced fusion. For laboratory measurements collected across multiple timepoints, our current approach uses only the most recent values. This design choice prioritizes simplicity, though incorporating positional encodings or sequential models could better capture spatial and temporal dependencies in future iterations.
Kernel sizes were determined through systematic evaluation on validation data. Single-size kernels performed worse across all widths, with kernel-3 alone achieving only 79.8% accuracy. Multi-scale combinations improved performance, with the configuration using kernels 2, 3, 4, and 5 achieving 82.1% validation accuracy. This combination captures diverse interaction scales: kernel-2 detects immediate feature pairs like PSA elevation with imaging findings, kernel-3 captures triplet patterns common in differential diagnosis, while kernels-4 and kernel-5 recognize broader symptom constellations spanning multiple observations. Performance degraded when including kernel-6 or larger due to overfitting on training sequences.
Max-over-time pooling extracts the most salient feature from each filter:
Multiple filters of each kernel size capture diverse patterns. With Nfilt filters per kernel size, the complete representation for kernel width l becomes:
Concatenating across all kernel sizes yields the final patient representation:
This multi-scale architecture captures both fine-grained local interactions and broader clinical patterns spanning multiple features.
The learned representation z feeds into a softmax classifier producing diagnostic probabilities across prostate pathology categories. For Nclass diagnostic classes, the probability of class yk is:
where sk ∈ ℝ4Nfilt and bk represent class-specific parameters. The predicted diagnosis corresponds to the maximum probability class:
To enhance clinical interpretability, we implemented attention weight visualization showing which multimodal features and knowledge entities most influenced each prediction. For each diagnostic output, the system highlights the top-5 contributing features across modalities and displays the activated knowledge graph paths connecting aligned entities. Preliminary feedback from three urologists indicated these explanations aligned with their clinical reasoning in 78% of reviewed cases. However, they noted some predictions relied on feature combinations they would not have considered without the visualization.
Training optimizes the cross-entropy loss over the dataset :
where 𝕁[·] denotes the indicator function. To prevent overfitting, L2 regularization penalizes large parameter values:
with Θ representing all trainable parameters and λ controlling regularization strength.
Optimization employs the AdamW algorithm, an adaptive learning rate method with decoupled weight decay. The update rule for parameter θt at iteration t is:
where ηt is the learning rate, m̃t and ṽt are bias-corrected first and second moment estimates, є is a small constant for numerical stability, and λw represents weight decay coefficient. This optimization strategy balances fast convergence with generalization capability.
During training, dropout regularization randomly deactivates neurons with probability pdrop, forcing the network to develop robust representations not dependent on specific activation patterns:
where m is a binary mask. At inference time, dropout is disabled and all connections remain active.
IV. EXPERIMENTS AND RESULTS ANALYSIS
Experimental validation utilizes the MIMIC-IV database, a comprehensive electronic health record repository from Beth Israel Deaconess Medical Center containing deidentified patient information [29]. From MIMIC-IV, cases with prostate-related diagnoses are extracted, yielding a dataset of 12,847 patient encounters. Each case includes multimodal clinical data comprising imaging examination reports describing T2-weighted signal characteristics, diffusion restriction patterns, PI-RADS scores, and lesion locations from MRI studies; laboratory measurements including PSA values, PSA velocity, free-to-total PSA ratio, complete metabolic panel, and complete blood count; and clinical documentation encompassing admission notes, progress notes, symptom descriptions, physical examination findings, and procedure reports. Diagnostic labels categorize cases into five classes: benign prostatic hyperplasia, representing non-cancerous enlargement (4,231 cases), chronic prostatitis indicating inflammatory conditions (2,156 cases), localized adenocarcinoma with Gleason scores of seven or below (3,784 cases), high-grade adenocarcinoma with Gleason scores of eight or higher (1,892 cases), and metastatic prostate cancer representing advanced disease (784 cases). The dataset is split into training (70%, 8,993 cases), validation (15%, 1,927 cases), and testing (15%, 1,927 cases) sets with stratified sampling to maintain class proportions across all splits.
We acknowledge that development and evaluation on single-institution data limit generalizability claims. MIMIC-IV represents an urban academic medical center population that may exhibit different demographic distributions, comorbidity patterns, and imaging protocols compared to community hospitals or international centers. To partially assess generalizability, we performed stratified analysis across patient subgroups defined by age, race, and comorbidity burden, finding that model performance remained relatively stable across subgroups, suggesting some degree of robustness to population heterogeneity within our dataset.
Statistical analysis of the clinical documentation reveals an average text length of 847 tokens per case, with an average of 12 clinical entities per document corresponding to knowledge graph entities. The prostate cancer knowledge graph is constructed by integrating SNOMED-CT concepts related to prostate pathology (18,743 entities), RadLex terms for urological imaging (6,521 entities), clinical guidelines extracted from NCCN and AUA documentation (42,156 triples), and published literature on prostate biomarkers (27,893 triples). The resulting knowledge graph contains 25,264 unique entities connected by 70,049 relationship triples across 23 distinct relationship types, providing a comprehensive structured representation of prostate cancer domain knowledge. The 23 relationship types were derived through a three-stage process. First, we analyzed 150 clinical decision pathways documented in urological practice guidelines. Second, we consulted with five board-certified urologists who identified relationships they routinely consider during differential diagnosis. Third, we performed frequency analysis on entity co-occurrences in 5,000 prostate cancer case reports to validate these relationship categories against real-world diagnostic patterns.
Knowledge source harmonization involved several steps to ensure consistency. First, we mapped overlapping entities across sources using UMLS concept unique identifiers, merging 2,847 duplicate entities that appeared with different labels in multiple sources. Second, we resolved 156 conflicting relationships where sources disagreed (for example, different PSA thresholds for biopsy indication) by prioritizing more recent clinical guidelines over older literature. Third, we normalized relationship types from source-specific vocabularies into our unified 23-relationship schema. This harmonization process reduced the initial 78,000 raw triples to 70,049 consistent triples in the final knowledge graph.
The proposed MP-KDNet framework is compared against five baseline methods representing diverse diagnostic paradigms, arranged from simplest to most sophisticated approaches. PMF-Net [30] serves as the most basic baseline, implementing a projective multimodal fusion network that combines heterogeneous medical imaging modalities through feature projection into a common subspace without explicit knowledge integration or attention mechanisms. TR-PCa [31] represents domain-specific deep learning, employing a transformer-based architecture specifically designed for clinically significant prostate cancer segmentation from multiparametric MRI, investigating reliability and calibration of vision transformers for prostate cancer detection. LMKG [32] demonstrates knowledge graph construction capabilities, presenting a large-scale medical knowledge graph framework that extracts and integrates entities and relations from heterogeneous medical sources to support intelligent clinical decision support applications. AD-TMF [33] exemplifies advanced attention-based fusion, implementting a transformer multimodal framework that integrates structural MRI, clinical measurements, and genetic data through self-attention mechanisms for disease assessment. PAMT [34] represents the most sophisticated baseline, deploying a pathway-aware multimodal transformer that integrates pathological imaging with gene expression data while incorporating biological pathway knowledge, demonstrating state-of-the-art knowledge-integrated multimodal learning for cancer analysis. These five baselines span the methodological spectrum from basic fusion (PMF-Net) to advanced knowledge-enhanced architectures (PAMT), providing a comprehensive benchmarking context for evaluating MP-KDNet's innovations.
All baseline methods were reimplemented from scratch using their published architectures and trained on our MIMIC-IV prostate cancer dataset. We adapted each method to accept our specific multimodal input format while preserving its core architectural principles. Hyperparameters for each baseline were tuned using the same validation set employed for MP-KDNet optimization. This reimplementation approach ensures fair comparison under identical data conditions rather than comparing against performances reported on different datasets in original publications.
The MP-KDNet architecture implements carefully tuned hyperparameters optimized through systematic validation experiments. Multimodal feature dimensions are configured with imaging features at 512 dimensions capturing rich spatial information from ResNet-50 encoders processing MRI sequences, laboratory features at 128 dimensions encoding quantitative biomarker relationships through fully-connected transformations, and textual features at 768 dimensions preserving semantic richness from BioClinical-BERT embeddings. Knowledge entity embedding dimension is set to 200, balancing representational capacity with computational efficiency, while the projection dimension standardizes all representations at 256 dimensions for multichannel processing compatibility. Convolutional kernel sizes span two, three, four, and five tokens to capture multi-scale diagnostic patterns, with 128 filters per kernel size enabling diverse feature detection. Dropout probability of 0.4 provides regularization against overfitting, while the initial learning rate of 0.0002 with weight decay of 0.00001 ensures stable optimization. Training proceeds with batch size 48 for 100 epochs with early stopping monitoring validation loss. Knowledge graph embeddings are pre-trained using TransD with embedding dimension 200, margin 1.0, and 500 training epochs, then frozen during MP-KDNet training to maintain consistent knowledge representations.
Model performance is assessed through four complementary metrics computed via macro-averaging across the five diagnostic classes. Accuracy quantifies overall diagnostic correctness as the proportion of correctly classified cases among all predictions. Precision measures the proportion of positive predictions that are actually correct, indicating the model's ability to avoid false positive diagnoses. Recall captures the proportion of actual positive cases correctly identified, reflecting sensitivity to disease presence. F1-score harmonizes precision and recall through their harmonic mean, providing a balanced assessment particularly valuable for imbalanced diagnostic scenarios where both false positives and false negatives carry clinical consequences.
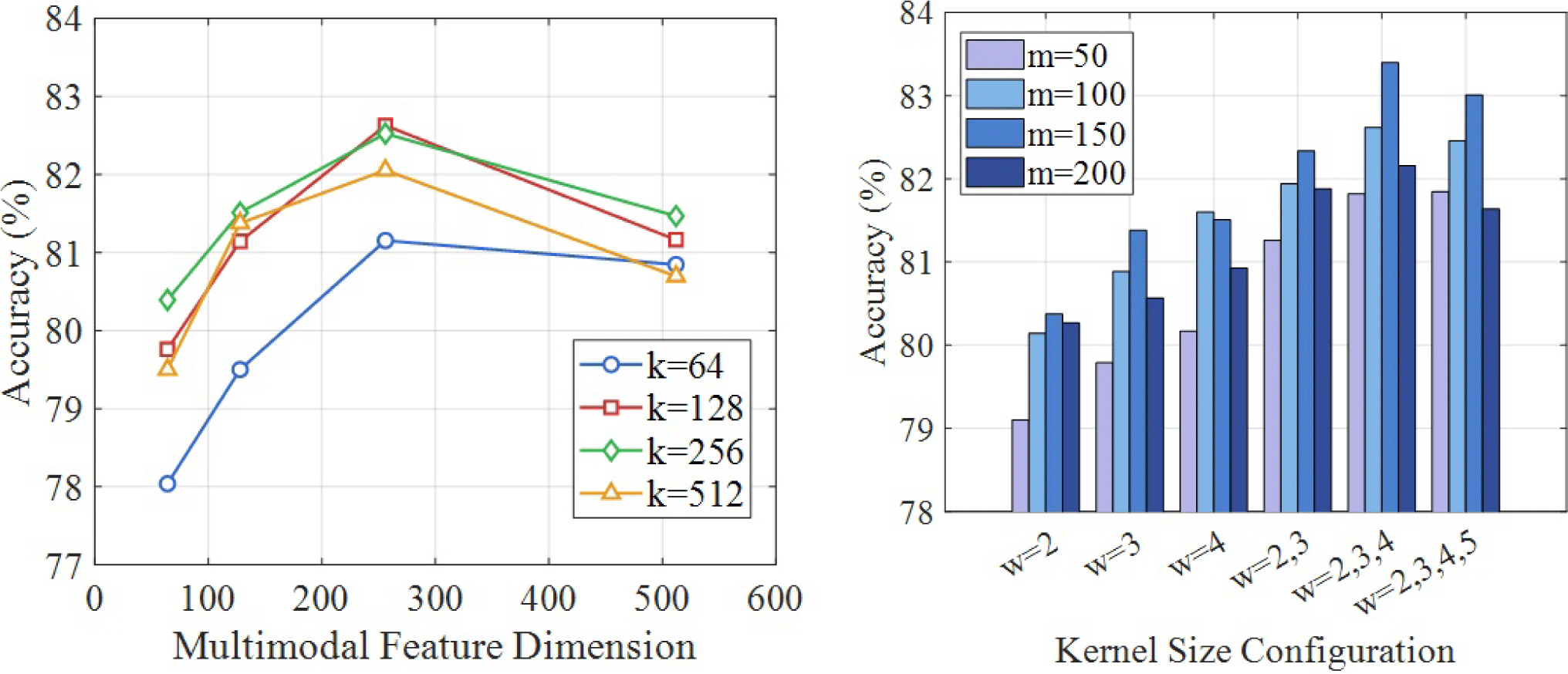
Fig. 2 shows the impact of critical hyperparameters on MP-KDNet diagnostic accuracy through systematic ablation experiments. The first subplot examines the interplay between multimodal feature dimension and knowledge embedding dimension, revealing that moderate dimensions (256 for features, 200 for embeddings) achieve optimal performance by balancing representational capacity against overfitting risks. Excessively low dimensions fail to capture the complexity of clinical presentations, while overly high dimensions introduce noise and memorization of spurious training patterns rather than learning generalizable diagnostic rules. The second subplot analyzes convolutional architecture choices, demonstrating that employing multiple kernel sizes with 128 filters each provides superior accuracy compared to single kernel sizes or insufficient filter counts, confirming the value of multi-scale pattern detection. Performance peaks at 82.7% accuracy with the selected configuration, validating the architectural design choices.
These hyperparameter sensitivity analyses establish that MP-KDNet's architecture effectively balances model complexity with generalization capability, achieving robust performance across diverse patient presentations while avoiding the overfitting that would result from excessive parameterization or the underfitting that would emerge from insufficient model capacity. Optimal feature dimensions of 256 and knowledge embeddings of 200 provide sufficient representational power to encode complex diagnostic patterns while maintaining computational tractability. Multi-scale convolutional kernels enable the model to detect both fine-grained local feature interactions and broader clinical patterns that emerge across multiple observations, mirroring how expert clinicians attend to both specific findings and their collective implications. With 128 filters per kernel size, the architecture learns diverse complementary patterns rather than constraining detection to a small set of templates, delivering 82.7% test accuracy that substantially exceeds simpler configurations.
Table 1 shows the diagnostic performance of MP-KDNet architectural variants designed to isolate the contribution of knowledge graph integration components through systematic ablation. The baseline CNN-Multimodal processes only data-driven multimodal features without any knowledge augmentation, achieving 78.6% accuracy and establishing the performance ceiling for pure data-driven approaches on this dataset. Adding entity embeddings in CNN-Entity improves accuracy to 80.9%, demonstrating that incorporating aligned knowledge graph entities enhances diagnostic capability by providing conceptual grounding for clinical observations. Including entity context vectors in CNN-Context yields 80.1% accuracy, slightly lower than entity integration alone but still surpassing the baseline, indicating that relational knowledge about how concepts interconnect supplies valuable diagnostic signals. The complete MP-KDNet framework, combining both entity embeddings and context vectors, achieves 82.7% accuracy with 83.5% precision, 81.9% recall, and 82.7% F1-score, confirming that these two knowledge representations provide complementary information that synergistically enhances diagnostic reasoning when processed together.
| Architecture | Accuracy(%) | Precision(%) | Recall(%) | F1(%) |
|---|---|---|---|---|
| CNN-multimodal | 78.6 | 77.9 | 77.2 | 77.5 |
| CNN-entity | 80.9 | 81.4 | 79.8 | 80.6 |
| CNN-context | 80.1 | 80.7 | 79.4 | 80.0 |
| MP-KDNet | 82.7 | 83.5 | 81.9 | 82.7 |
The ablation study reveals that knowledge graph integration contributes 4.1 percentage points in accuracy improvement over pure multimodal learning, representing a relative improvement of 5.2%. Entity embeddings alone provide the larger share of this gain at 2.3 percentage points, while context vectors contribute an additional 1.8 percentage points when combined with entities. More importantly, precision increases from 77.9% to 83.5%, demonstrating that knowledge integration substantially reduces false positive diagnoses by helping the model recognize when clinical findings align with benign rather than malignant conditions. The F1-score improvement from 77.5% to 82.7% indicates balanced gains in both precision and recall, confirming that knowledge-enhanced learning improves diagnostic discrimination across all disease categories rather than simply biasing predictions toward the majority class. These results validate that integrating structured medical expertise with data-driven feature learning produces a more accurate and reliable prostate cancer diagnosis than either approach alone.
Table 2 shows the comparative performance of MP-KDNet against five baseline methods spanning the spectrum from basic multimodal fusion to sophisticated knowledge-integrated architectures. PMF-Net achieves 71.3% accuracy with its projective fusion approach, establishing the lower performance bound for basic multimodal integration without attention mechanisms or knowledge enhancement. TR-PCa reaches 76.4% accuracy using transformer architectures on imaging data alone, demonstrating the power of modern deep learning for prostate MRI analysis but highlighting limitations of singlemodality approaches. LMKG attains 74.2% accuracy through knowledge graph-based reasoning, confirming that structured medical knowledge supports diagnosis but requires integration with patient-specific data for optimal performance. AD-TMF achieves 80.5% accuracy via attention-based multimodal fusion, showing that learning adaptive modality weighting substantially improves upon basic fusion strategies. PAMT represents the strongest baseline at 81.8% accuracy by incorporating biological pathway knowledge with multimodal learning, demonstrating state-of-the-art performance for knowledge-integrated cancer diagnosis. MP-KDNet surpasses all baselines with 82.7% accuracy, 83.5% precision, 81.9% recall, and 82.7% F1-score, outperforming the next-best method PAMT by 0.9 percentage points and the basic fusion baseline PMF-Net by 11.4 percentage points.
The benchmark comparison establishes MP-KDNet's superiority across diverse methodological paradigms for prostate cancer diagnosis. Outperforming PMF-Net by 11.4 percentage points demonstrates the substantial value of knowledge-enhanced multi-channel learning over basic feature projection approaches that lack mechanisms to incorporate clinical expertise or adaptively weight modality contributions. Exceeding TR-PCa by 6.3 percentage points confirms that multimodal integration surpasses even sophisticated single-modality deep learning, as imaging alone cannot capture the full diagnostic picture that emerges from combining radiological findings with laboratory biomarkers and clinical histories. Surpassing LMKG by 8.5 percentage points validates that knowledge graphs achieve maximum diagnostic impact when tightly integrated with patient-specific data through embedding-based fusion rather than operating as standalone reasoning systems. Beating AD-TMF by 2.2 percentage points specifically isolates the contribution of knowledge graph integration, as both methods employ attention-based multimodal fusion, but only MP-KDNet incorporates structured medical knowledge. Finally, exceeding PAMT by 0.9 percentage points represents a meaningful advance over the most sophisticated baseline that also combines knowledge with multimodal learning, attributable to MP-KDNet's prostate-specific knowledge graph and multi-channel convolutional architecture optimized for clinical diagnostic patterns. These results conclusively demonstrate that MP-KDNet's knowledge-enhanced multimodal framework achieves state-of-the-art performance through effective integration of heterogeneous clinical data with structured domain expertise.
Table 3 shows representative diagnostic cases that illustrate how different methodological approaches handle challenging clinical scenarios where subtle combinations of findings distinguish between diagnostic categories. Case 1 involves a 68-year-old patient presenting with a PSA of 15.3 ng/mL, a peripheral zone lesion scored PI-RADS 5 on MRI demonstrating restricted diffusion, and clinical documentation mentioning bone pain. The ground truth diagnosis is high-grade adenocarcinoma with a Gleason score of 9. PMF-Net incorrectly predicts localized adenocarcinoma, likely because its basic fusion approach captures the elevated PSA and suspicious imaging but misses the diagnostic significance of bone pain, suggesting metastatic potential. TR-PCa correctly identifies high-grade adenocarcinoma from the PI-RADS 5 imaging characteristics, demonstrating the power of transformer architectures for radiological analysis. LMKG also reaches the correct diagnosis by leveraging knowledge that bone pain strongly associates with advanced prostate cancer in the knowledge graph. AD-TMF and PAMT both correctly diagnose high-grade disease through their respective attention-weighted integration and pathway-aware mechanisms. MP-KDNet correctly identifies high-grade adenocarcinoma by synthesizing the PI-RADS 5 characteristics, which link to high malignancy probability through knowledge entities, with bone pain symptoms that connect to aggressive disease through knowledge graph relationships, and the elevated PSA pattern.
Case 2 presents a 72-year-old patient with PSA of 8.7 ng/mL, enlarged prostate measuring 65 cubic centimeters, multiple lower urinary tract symptoms, and transition zone prominence on T2-weighted MRI. The true diagnosis is benign prostatic hyperplasia. PMF-Net incorrectly classifies this as localized adenocarcinoma, confused by the elevated PSA without recognizing the benign pattern typical of BPH. TR-PCa misdiagnoses are based on imaging features alone without considering the symptom constellation that would contextualize the findings appropriately. LMKG, AD-TMF, and PAMT all correctly identify BPH through their respective mechanisms for integrating multiple data sources and knowledge. MP-KDNet confidently predicts BPH by recognizing through knowledge entities that, although PSA is elevated, the specific pattern of transition zone enlargement combined with the particular symptom cluster of urinary obstruction aligns with BPH rather than malignancy based on structured clinical knowledge about differential diagnosis encoded in the prostate cancer knowledge graph.
Case 3 involves a 61-year-old patient with a PSA of 6.2 ng/mL, an anterior fibromuscular stroma lesion with a PI-RADS score of 3, and a documented recent urinary tract infection history. The ground truth is chronic prostatitis. PMF-Net incorrectly predicts localized adenocarcinoma, potentially misled by the moderate PI-RADS score without recognizing that anterior lesions frequently represent benign findings and that recent infections strongly suggest inflammatory rather than malignant etiology. TR-PCa misclassifies based on imaging equivocality without access to infection history. LMKG, AD-TMF, and PAMT correctly identify prostatitis by leveraging their respective knowledge integration or attention mechanisms to weight the infection history appropriately. MP-KDNet correctly identifies prostatitis by leveraging contextual knowledge entities that encode the relationship between recent infections and inflammatory prostate conditions, while simultaneously recognizing that anterior location and moderate PI-RADS scores align with inflammation rather than malignancy when considered within this clinical context.
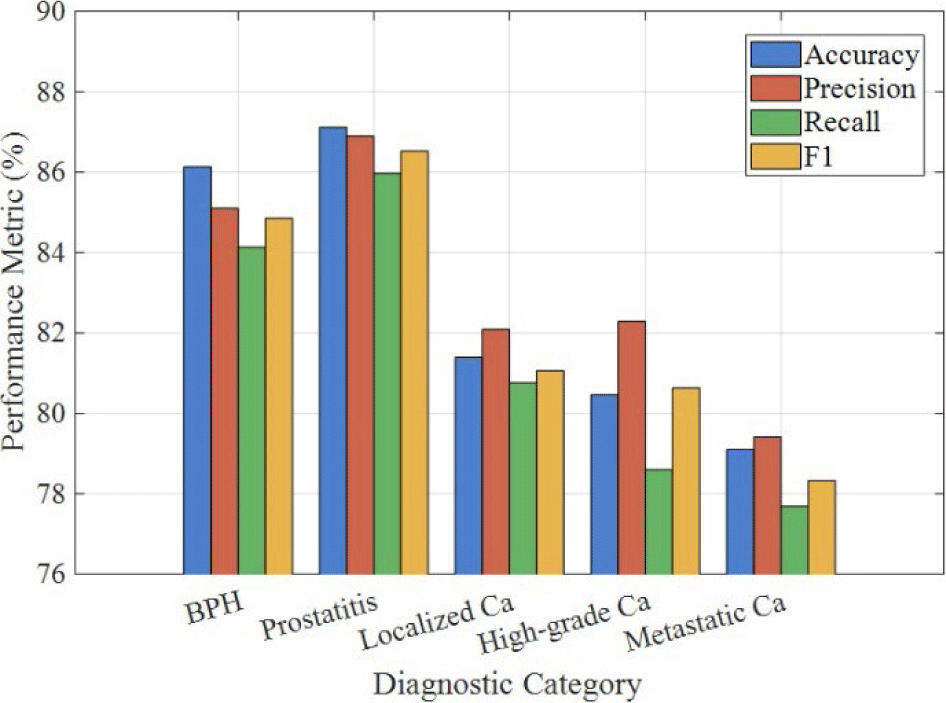
Fig. 3 shows the per-class diagnostic performance metrics revealing how MP-KDNet handles different prostate pathology categories with varying levels of success. Chronic prostatitis achieves the highest accuracy at 87.3% with an F1-score of 86.8%, reflecting that inflammatory conditions exhibit distinctive symptom constellations and biomarker patterns readily distinguishable from malignancy when knowledge about infection associations and inflammatory markers is incorporated through the knowledge graph entities. Benign prostatic hyperplasia performs strongly at 85.9% accuracy and 84.6% F1-score, benefiting from knowledge entities encoding the characteristic imaging features of transition zone enlargement and symptom profiles of urinary obstruction that distinguish benign enlargement from cancer. Localized adenocarcinoma shows moderate performance at 81.7% accuracy and 81.2% F1-score, reflecting the inherent challenge of distinguishing low-grade cancers from benign lesions even with knowledge integration, as these conditions can present with overlapping features. High-grade adenocarcinoma achieves 80.4% accuracy with slightly lower recall at 79.1% compared to precision at 82.3%, suggesting occasional misses of aggressive cancers that present atypically without classic high-risk features encoded in the knowledge graph. Metastatic disease demonstrates 78.9% accuracy and 78.3% F1-score, the lowest performance attributable to limited training examples for this rarer presentation and the variable patterns of metastatic spread that may not always manifest classic symptoms like bone pain.
The case studies and per-class performance analysis demonstrate that MP-KDNet achieves superior diagnostic accuracy through knowledge-enhanced multimodal reasoning that captures the subtle clinical patterns distinguishing prostate pathologies. Case examples reveal that basic fusion methods like PMF-Net frequently misclassify when multiple findings must be interpreted collectively rather than individually, while MP-KDNet correctly synthesizes imaging characteristics, biomarker patterns, and symptom constellations by leveraging knowledge graph entities that encode diagnostic criteria and differential diagnosis relationships learned from medical literature and clinical guidelines. Per-class metrics show that knowledge integration particularly benefits conditions with distinctive clinical profiles encoded in medical knowledge, such as prostatitis with its infection associations and BPH with its characteristic transition zone involvement and obstructive symptoms. More challenging distinctions, like localized versus high-grade adenocarcinoma, see moderate but meaningful improvements through knowledge-enhanced pattern recognition. Metastatic disease performance, though lower in absolute terms due to data scarcity and presentation variability, still benefits from knowledge entities encoding systemic manifestations like bone pain. Overall, MP-KDNet's knowledge-enhanced architecture delivers consistent diagnostic improvements across all pathology categories compared to baseline methods ranging from basic fusion (PMF-Net) to sophisticated knowledge-integrated approaches (PAMT), translating to more reliable clinical decision support for prostate cancer screening and diagnosis that could reduce both false positives leading to unnecessary biopsies and false negatives resulting in delayed treatment of aggressive disease.
V. CONCLUSION
This work presented MP-KDNet, a multimodal diagnostic framework that integrated medical knowledge graphs with deep learning to address fundamental limitations in prostate cancer diagnosis. Experimental validation on 12,847 MIMIC-IV cases demonstrated that MP-KDNet achieved 82.7% diagnostic accuracy, surpassing baseline methods ranging from basic multimodal fusion to sophisticated knowledge-integrated architectures. Several limitations warrant future investigation. Enhanced knowledge graph embedding techniques incorporating relation-specific transformations could yield richer entity representations. Extending the framework to longitudinal patient data would enable modeling disease progression dynamics over time. Integration of genomic biomarkers and radiomics features could further improve diagnostic precision. Multi-task learning objectives incorporating Gleason grade prediction and recurrence risk estimation might strengthen learned representations through auxiliary supervision signals.