I. INTRODUCTION
With the advancement in data capturing technologies, the volume of data is growing exponentially year by year. Traditional methods fail to provide an efficient mechanism for analysing and extracting useful information from such a large volume of data. Machine learning has appeared to be the perfect solution to this problem. The ability of a machine learning system to draw useful information from complex multi-dimensional data makes its usage ubiquitous i.e. in Research and Education, Transportation, Manufacturing, Healthcare, Military, etc.
Healthcare industry makes extensive use of machine learning algorithms, especially in the field of medical diagnosis and drug discovery [1]. In medical diagnosis, supervised machine learning algorithms are used to first analyse the dataset and extract the hidden information within it, thereafter this knowledge is used for diagnosing any previously unseen or future cases [2][3].
The nature of the input data plays a significant role in determining the performance of a machine learning algorithm. There are algorithms which work exceptionally well with only normalized data [4], but some algorithms work equally well with both normalized and un-normalized data. Thus the choice of the algorithm plays a very important role in determining the performance of the resulting system.
This paper illustrates a comparative analysis of performance of 4 machine learning algorithms i.e. LDA, Naive Bayes (NB), k-Nearest Neighbours (KNN) and Support Vector Machines (SVM) on the basis of their classification accuracy. The whole paper is divided into 7 sections, i.e. introduction, literature review, data pre-processing, methodology, results and discussion, conclusion and finally the future scope. This section gives brief introduction about the field and its area of application, next section gives a brief review of the corresponding literature, followed by data preprocessing, methodology & experimentation, results and discussion, conclusion and the future scope.
II. LITERATURE REVIEW
Fung et al. [5] proposed a linear programming based SVM model which selects the important voxels and also provides the most important areas for classification. The authors implemented their model on data from different European institutes. The authors obtained a sensitivity of 84.4% and a specificity of 90.9% which was then compared with the results obtained from Fischer linear discriminant (FLD) classifier and Statistical parametric mapping (SPM). The given approach outperformed human experts and both FLD and SPM. Gorriz et al. [6] created an automatic system for diagnosing Alzheimer’s disease in its early stages. They searched for discriminant Region of interests (ROIs) with different shapes as a combination of voxels in the masked brain volume. Each ROI was used for training and testing for SVM classifier which created an ensemble of classification data. The authors used pasting vote technique to aggregate this data using two different sum functions. It was observed that the size of ROIs was more significant for the performance of the classifier as compared to their shape. The pasting-vote function which aggregated the weighted summation of votes having relevant information from ROIs gave the best accuracy. Authors obtained an accuracy of 88.6% using this approach. Horn et al. [7] performed differential diagnosis of Alzheimer’s disease (AD) and Fronto- Temporal Dementia (TD) using various linear and non-linear classifiers on Single photon emission computed tomography (SPECT) data obtained from multiple hospitals. A total of 116 attributes were obtained as ROI from the SPECT images of 82 AD and 91 FTD patients. The classifiers selected for the experiment were a linear regression (LR), Linear discriminant analysis (LDA), SVM, KNN, Multi-layer perceptron (MLP) and K-logistic Partial least squared (PLS). These classifiers were used in different combinations and their performance in terms of classification accuracies was compared with each other and with 4 physicians. The best performance was obtained when SVM and PLS was combined with KNN. This combination achieved a classification accuracy of 88% which was higher than that of the physicians (accuracy values ranged from 65% to 72%).
López et al. [8] proposed an automatic diagnostic system for Alzheimer’s disease using SVM, Principal component analysis (PCA) and LDA based upon SPECT images collected from 91 patients. Authors first extracted the features from the given images using LDA, thereafter the significant features were selected using K-PCA. The data obtained was used for the training of SVM classifier which gave a classification accuracy of 92.31%. The given system outperformed the traditional approach i.e. voxels-as-features (VAF) which gave a classification accuracy of 80.22%. Huang et al. [9] proposed an automated method for diagnosis of Alzheimer’s where they used the cortical thickness from brain Magnetic resonance imaging (MRI) images as features for the classification process. Authors created Degenerate AdaBoost featuring an AdaBoost method based upon SVM. The authors compared the performance of the proposed system with the traditional classifiers i.e. SVM, KNN, LDA and Gaussian mixture model (GMM) and found that the proposed system outperformed all other classifiers with an accuracy of 84.38%. Alam et al. [10] combined the features extracted from structural MRI (sMRI) images obtained from Alzheimer’s disease neuroimaging initiative (ADNI) with those of Mini-mental state examination (MSME) scores of the given patients for differential diagnosis of AD and Mild cognitive impairment (MCI) from Healthy controls. The authors first performed two sample t-test for selecting a subset of the features. The selected subset is then fed to the kernel PCA (KPCA) for projecting the obtained data onto reduced PCC at higher dimensional space for increasing the linear separability. These kernel PCA coefficients were then projected into linear discriminant space using LDA. Finally a multi-kernel SVM (MKSVM) was used to perform the classification based on this data. For AD vs Healthy control classification, the chosen model gave an accuracy of 93.85% whereas for MCI vs HC and MCI vs AD the proposed method gave accuracies of 86.4% and 75.12% respectively.
III. DATA PRE-PROCESSING
For the purpose of this study, Alzheimer’s dataset from kaggle.com was taken. The dataset consisted of 373 records and a total of 14 independent attributes in the original dataset namely Subject_ID, MR_Delay, MRI_ID, Visit, M_F, Age, Hand, EDUC, MMSE, SES, nWBV, CDR, eTIV, and ASF. The attribute values represented clinical and other test results obtained from the longitudinal study of patients under consideration for the respective study. After initial screening, Subject_ID, MRI_ID, MR_Delay, Visit, and Hand were removed from the given dataset as these had no significant information for the classifier. Hence the dataset was left with only 9 predictor attributes after the initial screening phase. Group was the dependent variable which represented 3 classes i.e. Converted = 37, Non-Demented=190, Demented=146 and instances respectively. Before applying any pre-processing, all the attribute values were first transformed into numeric values by performing required conversions. Also, the dataset had some missing values for SES and MMSE. Local Mean was applied on the given columns to impute the missing values.
After imputation, the attribute values were normalized by applying Min-Max Normalization process given by,
where v′ = normalized value, v = original value of the attribute, vmn = minimum value, vmx = maximum value respectively for the given attribute.
IV. METHODOLOGY & EXPERIMENTATION
Two different versions of the given dataset were used for performing the experiments:
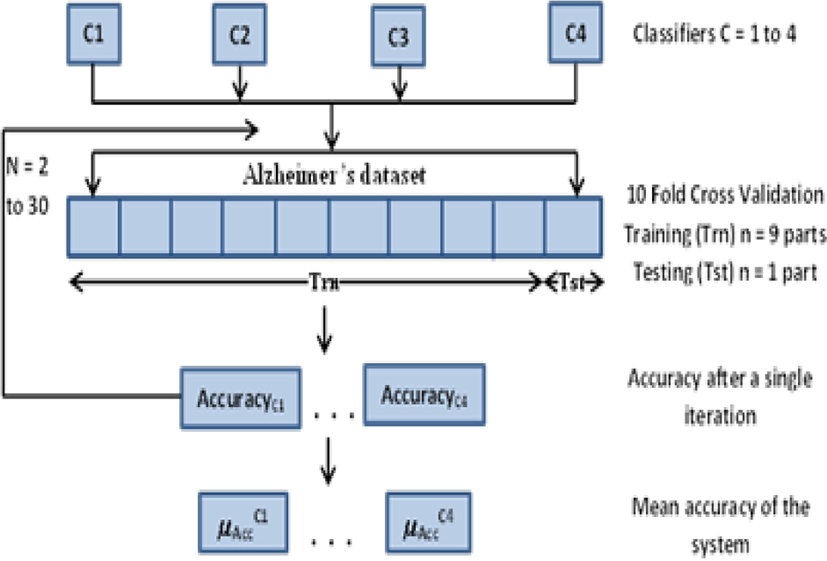
This paper performed a comparative analysis of LDA, NB, KNN and SVM on Alzheimer’s dataset. These algorithms have been frequently used in the past for building up of Computer based Diagnostic Systems (CDS) [11][12], that’s why they were included in this study. Fig. 1 shows the proposed architecture.
The complete experiment was implemented in python 2.7 using jupyter notebook. The given classifiers were run on both normalized and un-normalized data from the Alzheimer’s dataset obtained from kaggle.com. Accuracy was chosen as the performance metrics.
Accuracy is the ratio of correctly classified cases to that of the total no of cases under consideration and is calculated as,
where TP = True Positive, i.e. cases that are correctly classified as positive by the classifier.
TN = True Negative, i.e. cases that are correctly classified as negative by the classifier.
FP = False Positive, i.e. cases that are negative but classified as positive by the classifier.
FN = False Negative, i.e. cases that are positive but classified as negative by the classifier.
For both normalized and un-normalized data, the experiment was carried out 30 times to obtain consistent and reliable results. 10 fold cross-validation was used for cross-checking the validity of the obtained accuracy values. For each iteration, the complete dataset was divided into 10 folds. Out of these 10 folds, 9 were used for training and 1 fold was used for testing in such a manner that all the folds must be used for testing at-least once. This type of setup is known as 10 fold Cross-Validation or K-fold Cross-Validation in general. In each iteration an accuracy score was obtained for each classifier. The mean of the accuracies of each classifier for all the 30 iterations was taken as the final value of classification accuracies for the respective classifiers. The results obtained are discussed briefly in the next section.
V. RESULTS & DISCUSSION
Table1 lists the findings of this experiment. It shows the accuracy values for the given classifiers on both un-normalized and normalized data. It can be seen that for both normalized and un-normalized data, LDA gives the best accuracy i.e. 89.83%, whereas KNN has the least accuracy i.e. 46.87% and 82.80% w.r.t un-normalized and normalized data from the given dataset.
| Classifier | Accuracy (%age) | %age improvement | |
|---|---|---|---|
| Un-Normalized | Normalized | ||
| LDA | 89.83 | 89.83 | 0% |
| KNN | 46.87 | 82.80 | 76.66% |
| NB | 88.19 | 88.19 | 0% |
| SVM | 53.40 | 88.75 | 66.20% |
Another very important observation from Table 1 is the difference in the classifier accuracies on un-normalized and normalized data. It is evident from Table 1 that KNN and SVM do not perform well on un-normalized data but their performance improves significantly when applied on normalized data. This is attributed to the fact that KNN and SVM perform no internal normalization before classification process and give more importance to higher weighted attributes. This results in decrease in overall accuracy as it gives more importance to some attributes (due to higher values) and less importance to others (with smaller values). Whereas, LDA and NB perform equally well on both normalized and un-normalized data. This is because LDA and NB perform internal normalization on the given data before performing classification and also NB assumes attributes to be independent of each other. These facts can be inferred from Fig. 2 and Fig. 3 which represent the performance of the classifiers on both un-normalized (UND) and normalized data (ND) and the percent improvement in accuracy from un-normalized to normalized data respectively.
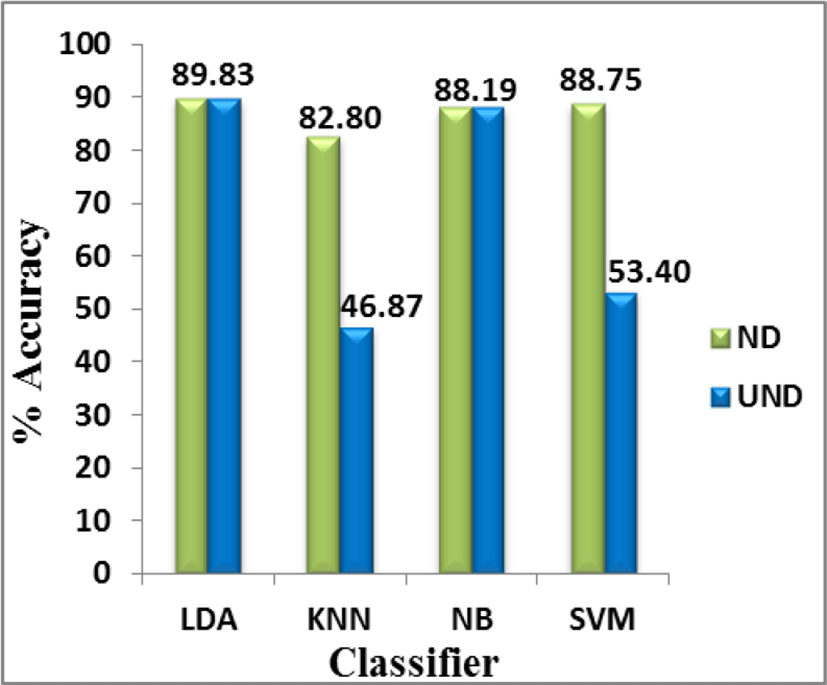
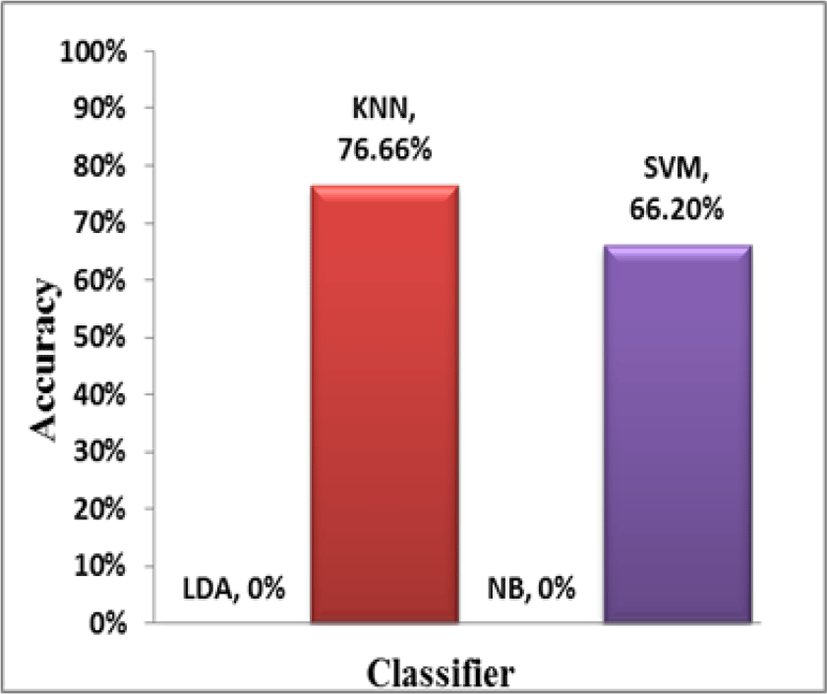
It can be seen that LDA and NB show no improvement in accuracy when migrated from un-normalized to normalized data, whereas KNN and SVM show 76.66% and 66.20% improvement in accuracies respectively when migrated from un-normalized data.
Authors compared their work with the work done by different authors in the similar domain or research problem. From Table 2, it can be seen that the accuracy of the proposed model is comparable to that of [13], however it is less than [14] and [15], the reason for this is that the current research listed a 3 class problem with imbalance in the classes as compared to the 2 class problem of others. Further the main focus of this research is to check the behaviour of different algorithms on both normalized and un-normalized data. Out of the different work shown in Table 2, only [15] compared the results of the classifier on both noisy and non-noisy data in which the performance of the best classifier i.e. Recursive feature selection based SVM (RFS-SVM) improved form 82.56% to 98.92% i.e. an improvement of about 20%. However, the current research showed an improvement of about 76.66% (46.87% to 82.80%) and 66.20% (53.40% to 88.75%) for KNN and SVM respectively as is evident from Table 2.
| Authors | Classifiers | Diseases | Classes (2C/3C) | Accuracy/f1 Score |
|---|---|---|---|---|
| [13] | AdaBoost | Alzheimer’s | 2 Class | 79.6 |
| MCI | 90.1 | |||
| [14] | k-NN | Breast Cancer | 2 Class | 94.1 |
| [15] | RFS-SVM | Diabetes | 2 Class | 98.92 |
| [16] | Random Forest | Diabetes | 2 Class | 89.63 |
| Our method | LDA | Alzheimer's | 3 Class | 89.83 |
VI. CONCLUSION
From the given experiment, it is concluded that LDA shows the best performance on the given Alzheimer’s dataset for a 3 class problem. Further, it is also concluded that LDA and NB perform equally well on both normalized and un-normalized data. However KNN and SVM show poor performance on un-normalized data, but their performance improves by a significant level when applied on normalized data.